Electron Microscopy DNA/Single Molecules
Unit info
The core Transmission Electron Microscopy (TEM) facility single molecules of IFOM specializes in the visualization of nucleic acids (DNA) and proteins using high-resolution and three-dimensional (TEM) techniques. DNA and proteins are the fundamental components of the chromosomes that are the cellular structures devoted to preserve and transmit the genetic information in the living organisms. Although the atomic ultra-structure of the DNA molecule is known from more than 70 years, at present, the in vivo DNA structures underlaying essential cellular processes to maintain and transmit the chromosomes to the daughter cells like DNA replication, DNA repair, DNA transcription and DNA recombination processes are almost completely unknown. Also specialized DNA-based structural elements of the chromosomes, including centromeres, telomeres, repetitive DNA sequences (satellite DNA) and extrachromosomal DNA elements like DNA circles and mitochondrial DNA have uncertain in vivo DNA structures. Proper regulation of the duplication and maintenance of the chromosomes through the above-mentioned DNA processes and their balanced interactions are crucial for maintaining genome stability thus preventing the insurgence of mutations and chromosomal alterations, which are hallmark of cancer.
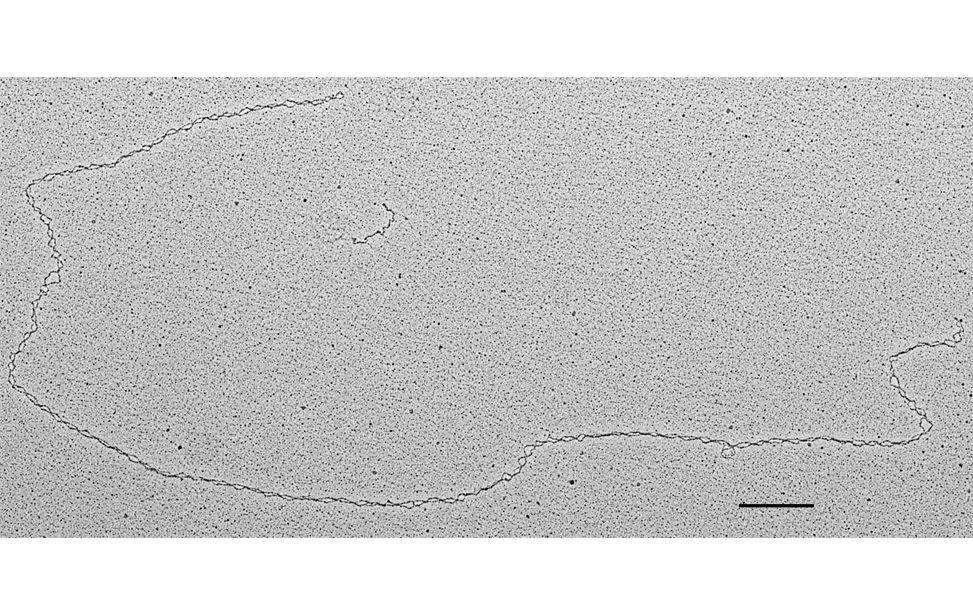
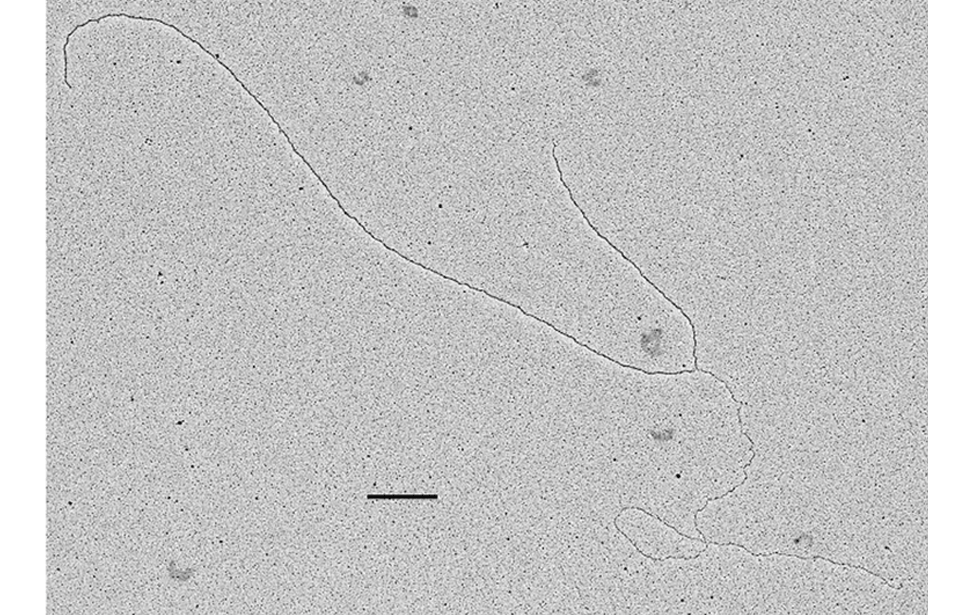
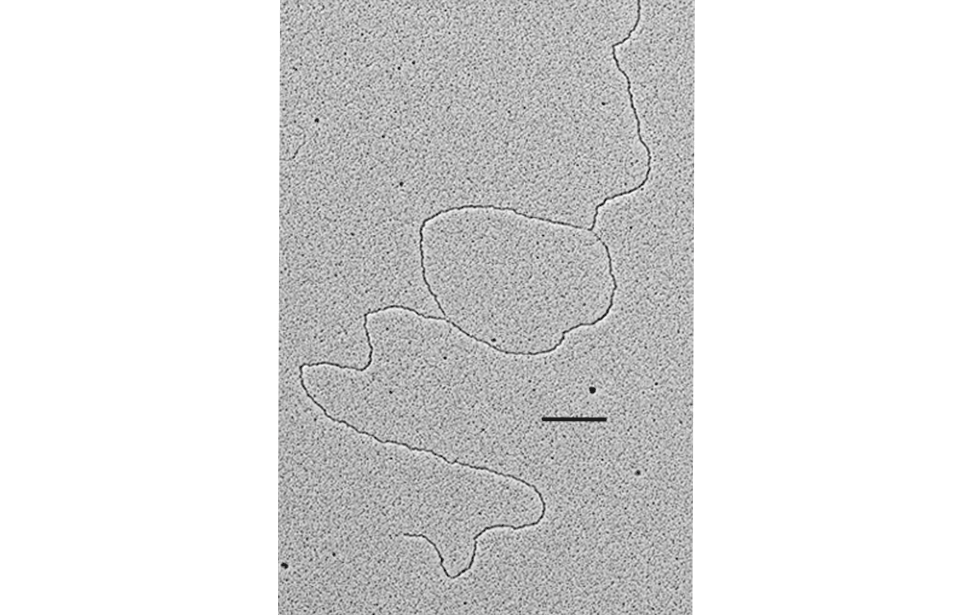
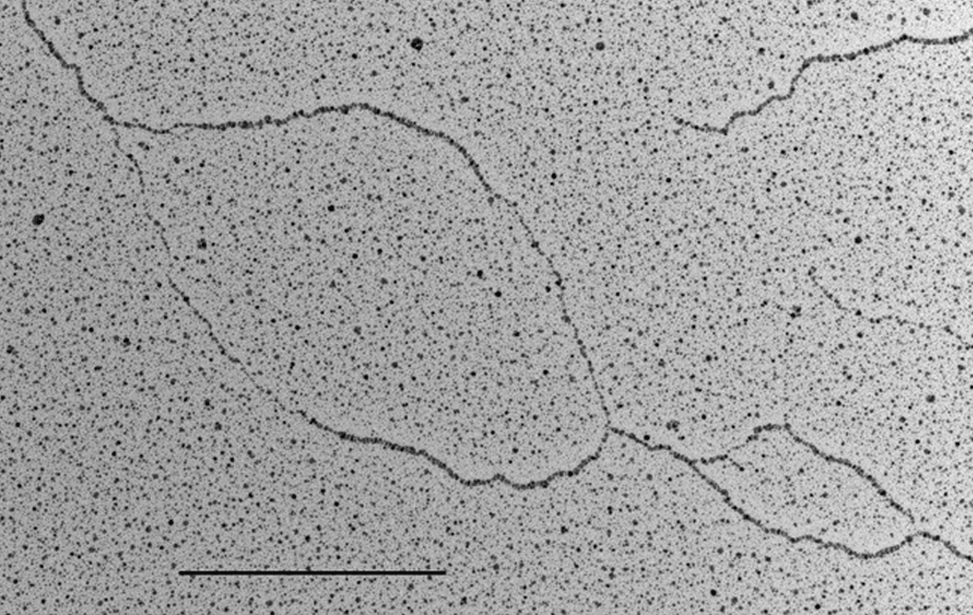
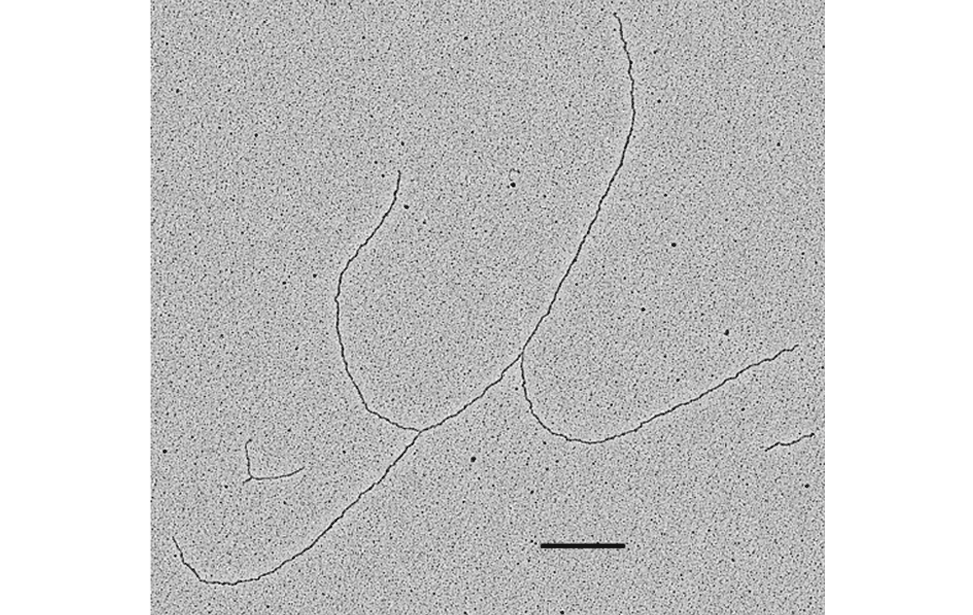
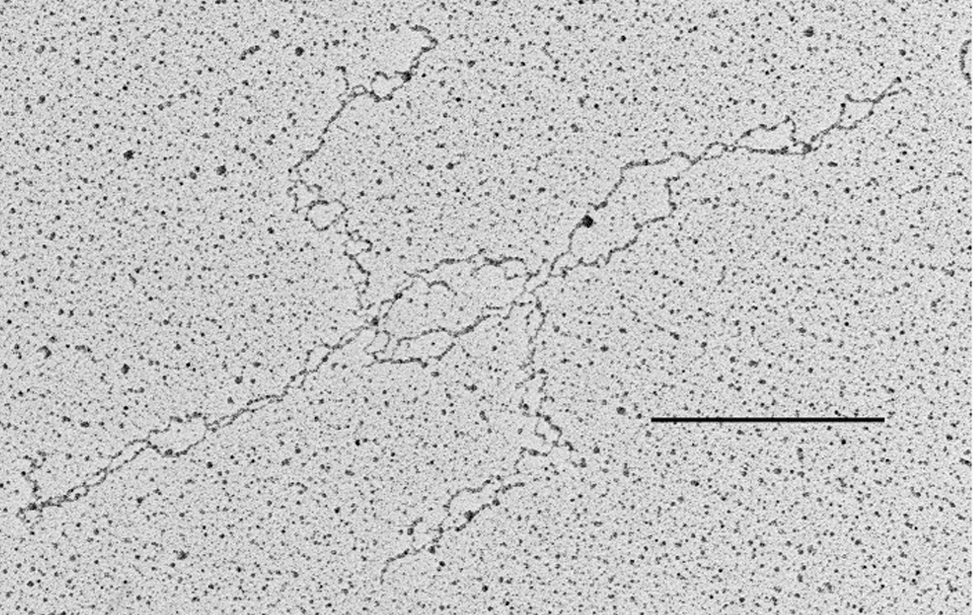
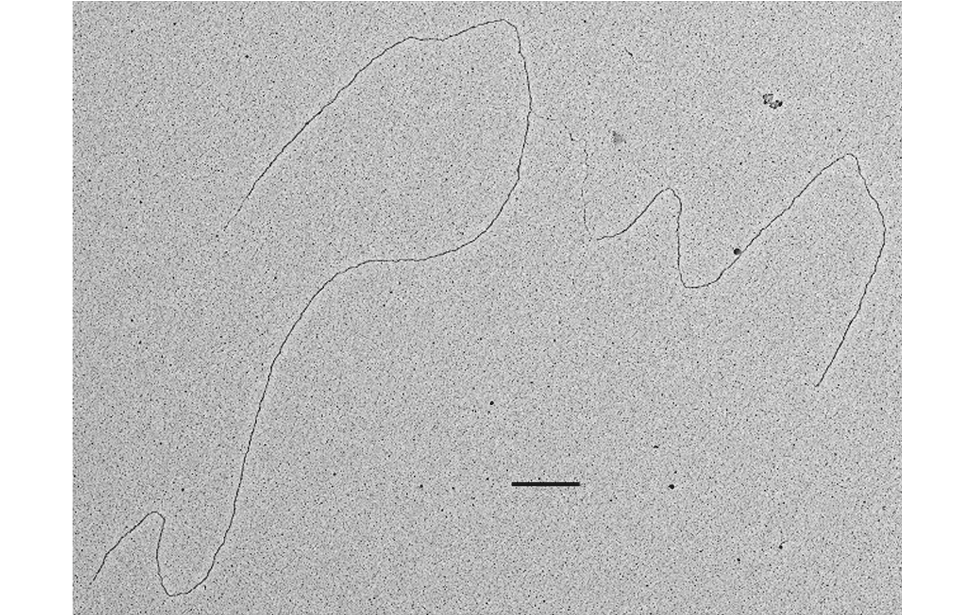
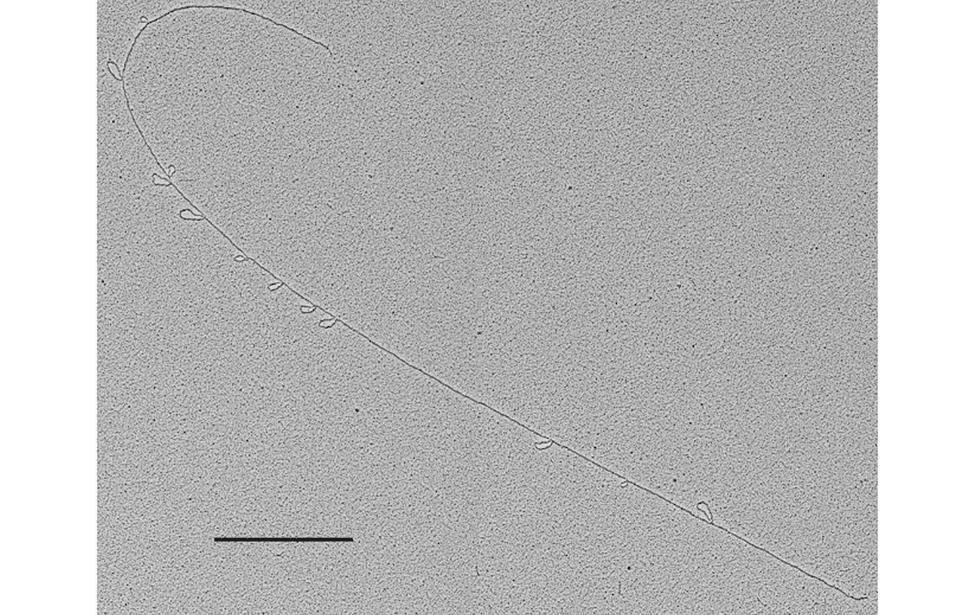
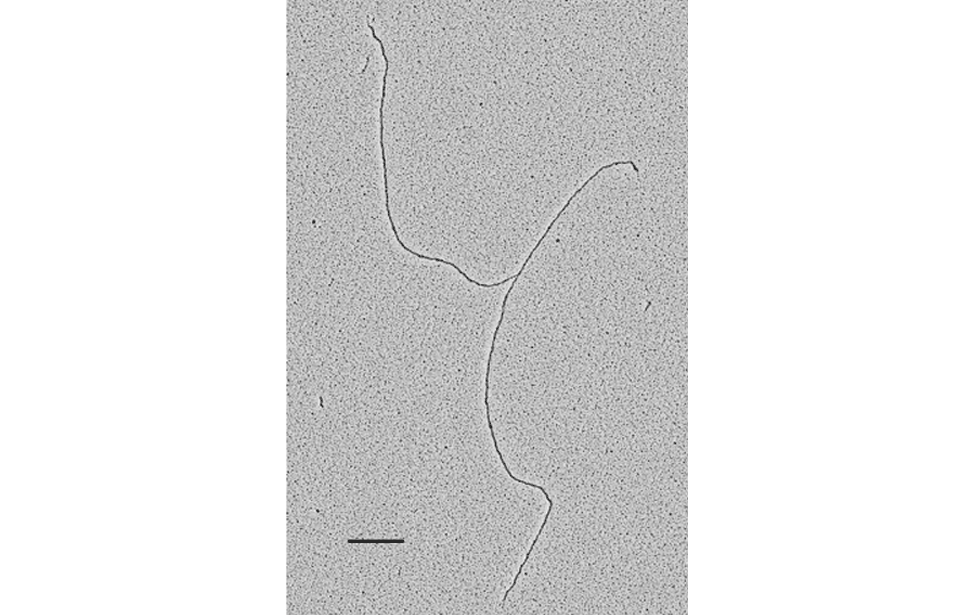
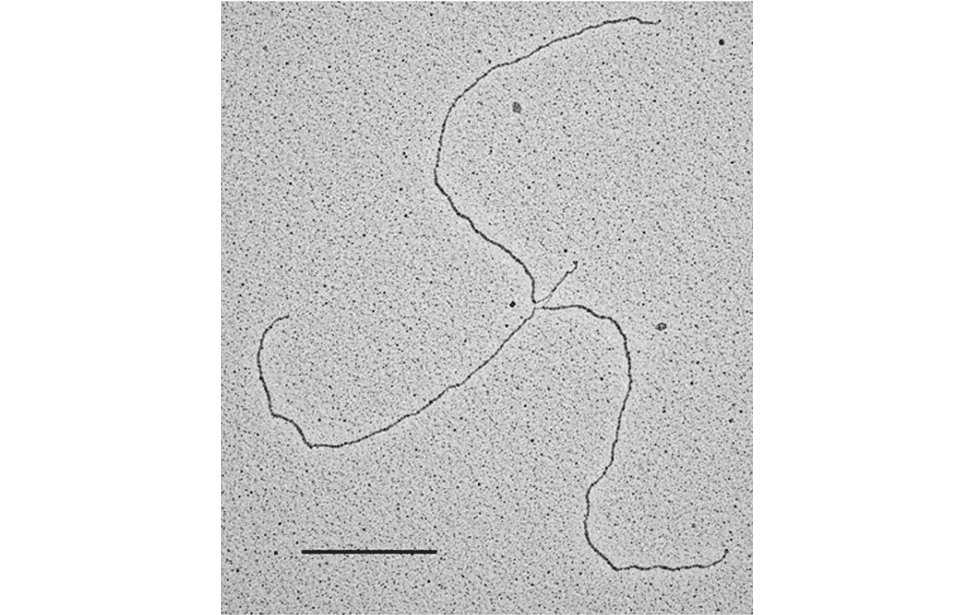
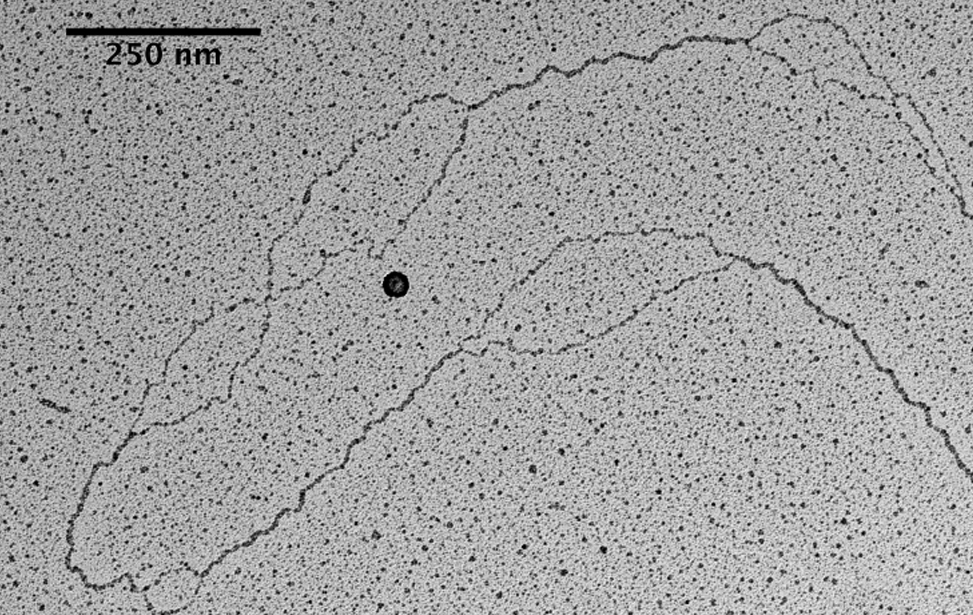
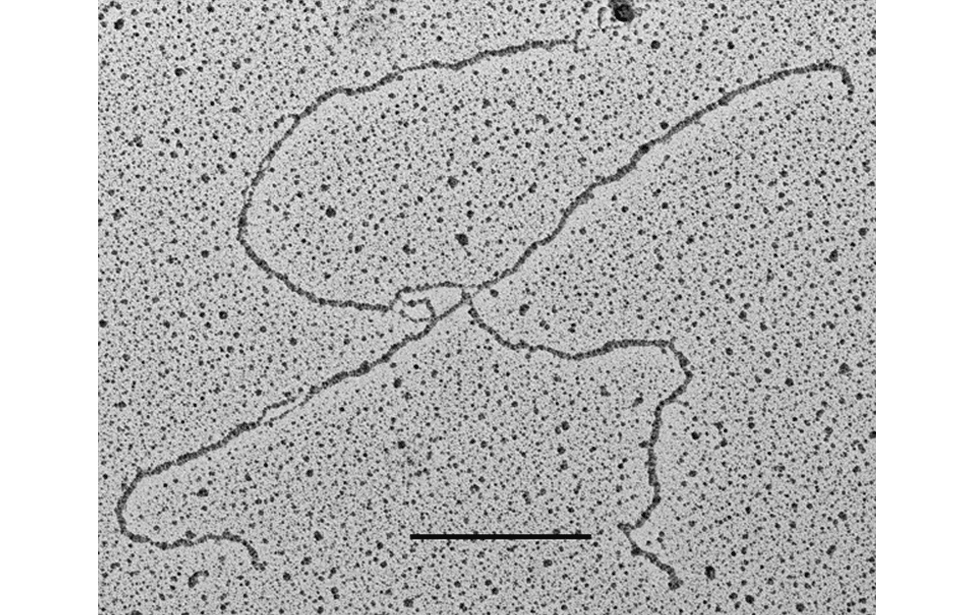
The facility uses the rotary shadowing technique coupled to TEM to inspect the in vivo DNA structures of the chromosomes once they have been properly purified and enriched from the large majority of the not-informative linear DNA. To visualize and understand the DNA structures underlying fundamental cellular processes involved in the metabolism of the chromosomes (at a molecular level) is a key approach for the understanding of the cellular mechanisms that govern how the genetic material is preserved in the normal cells and how the malfunction of these processes can lead to the insurgence of genome instability and to the acquired capability of the cancer cell to divide in an uncontrolled way.
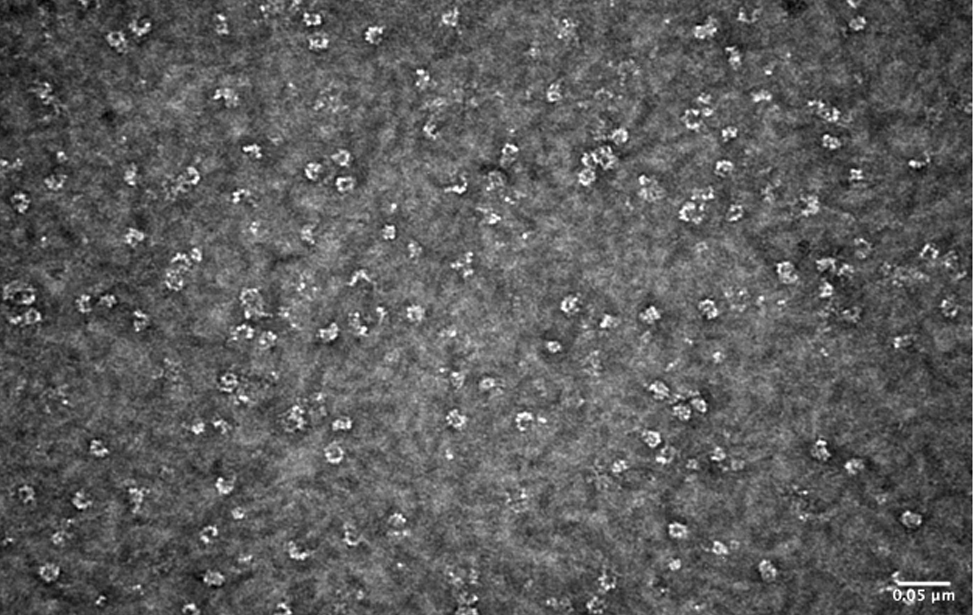
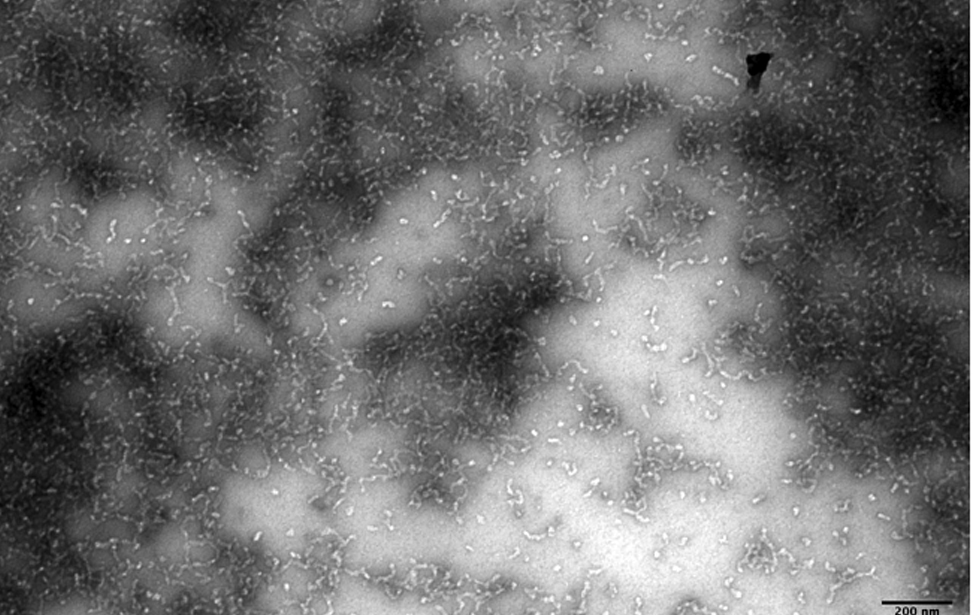
The facility also focuses on the study of the structure of the proteins using negative staining and rotary shadowing single molecule thecniques. Disease and cancer induced protein protofibrils, fibrils, protein misfolding and the structure of DNA-protein complexes are subjects of study of the facility. It is of key importance to understand the biology of the cancer cell to understand the molecular mechanisms through which the complex DNA and DNA-protein-mediated molecular assemblies preserve the integrity of the genetic material and its proper transmission to the daughter cells.
The facility actively trains researchers, enabling them to apply complex single molecules techniques in their research projects. By fostering collaboration within IFOM, the facility contributes to the ongoing development of new methods for studying the in vivo DNA structures the supramolecular DNA-protein mediated complexes either in physiological conditions or in the disease.
The Electron Microscopy Single Molecule Core Facility supports a variety of research projects by offering high-resolution ad three-dimensional imaging techniques to visualize nucleic acids, proteins and protein-nucleic acid complexes with high resolution. Key applications include:
Structural Analysis of DNA and RNA structures
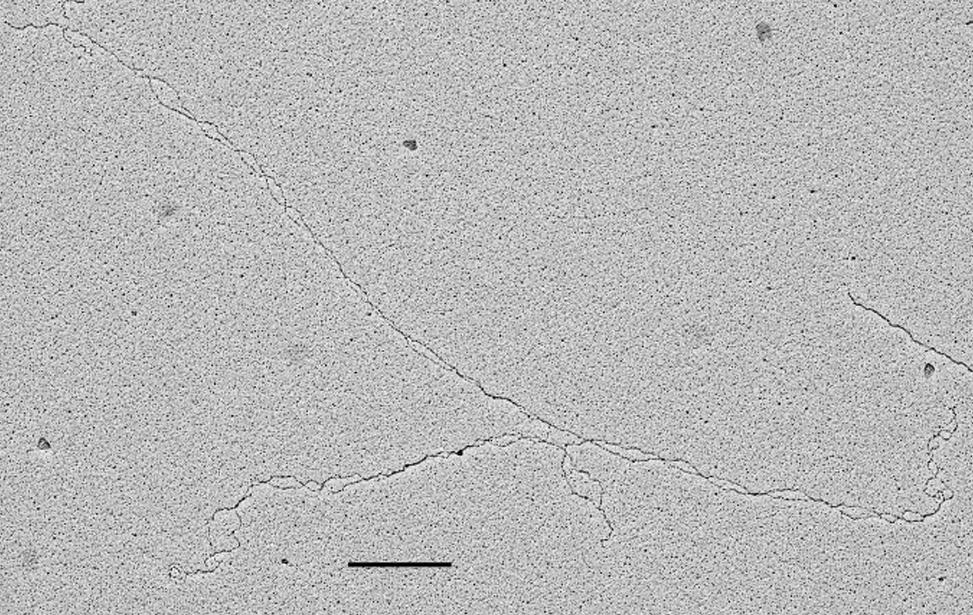
- Visualization of the in vivo structure of chromosomal DNA and DNA-RNA structures involved in DNA replication, DNA repair, DNA recombination and DNA transcription.
- Investigation of the in vivo structure of structural DNA elements of the chromosomes like repetitive DNA sequences, including centromeres, telomeres, and satellite DNA .
- Structural characterization of episomal DNA elements like extra-chromosomal DNA circles and mitochondrial DNA replication intermediates.
Genome Instability and Cancer Research
- Identification of cancer-associated nucleic acid structures.
- Ultra-structural analysis of DNA, RNA, and DNA-RNA intermediates for potential diagnostic applications.
Proteins and DNA-Protein Complexes
- Visualization of the structure of the proteins and DNA-protein complexes (through negative staining or rotary shadowing). Structure of the protein protofibrils, fibrils and supramolecular protein assembly in normal cells and disease derived cells.
- 2D class averaging and low-resolution 3D reconstruction of protein structure (from negative staining datasets) using the open source CryoSparc (https://cryosparc.com/).
Knowledge Transfer & Network
- Training IFOM researchers in complex single molecule electron microscopy techniques.
- Building a network of skilled users that can apply advanced electron microscopy techniques, methodological innovation.
Future implementations
It is programmed that, in the next future, the electron microscopy facility DNA/single molecules will acquire an automatic plunge freezer instrument so that the complete pipe-line (i.e. automatic plunge freezer, TALOS F200C G2 transmission microscope, the CetaS/F high sensitive, high S/N camera, the Gatan 626 side entry cryo-transfer sample holder, the EPU and the cryosparc 3D-reconstruction software platforms) to analyse the 3D structure of the single bio-molecules in their native states in vitreous ice (without staining with heavy metals), will be owned by the IFOM facility and will become available for the IFOM/AIRC research scientists
The facility collaborates closely with IFOM research groups, including those led by Dana Branzei, Vincenzo Costanzo, Fabrizio d’Adda di Fagagna, Ylli Doksani, Marco Foiani, Angela Bachi, and Simona Polo. These partnerships contribute to advancing research on genome integrity and cancer biology, key areas within Athena, IFOM’s ambitious strategic research plan.
La facility EMSM è dotata di strumentazione avanzata per lo studio dell’ultrastruttura e delle interazioni molecolari tramite microscopia elettronica. Include:
- ThermoFisher Transmission electron microscope TALOS F200C G2
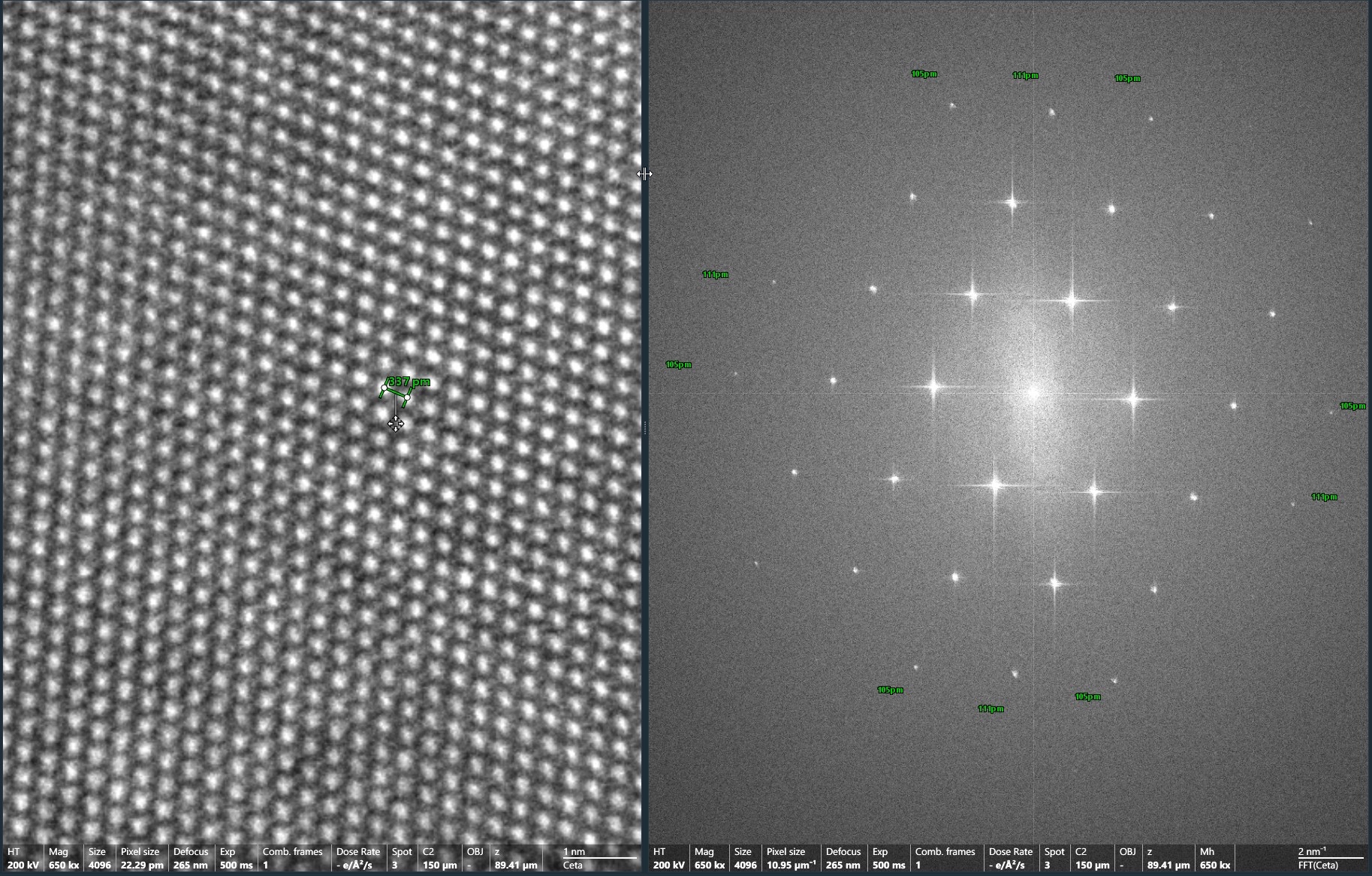
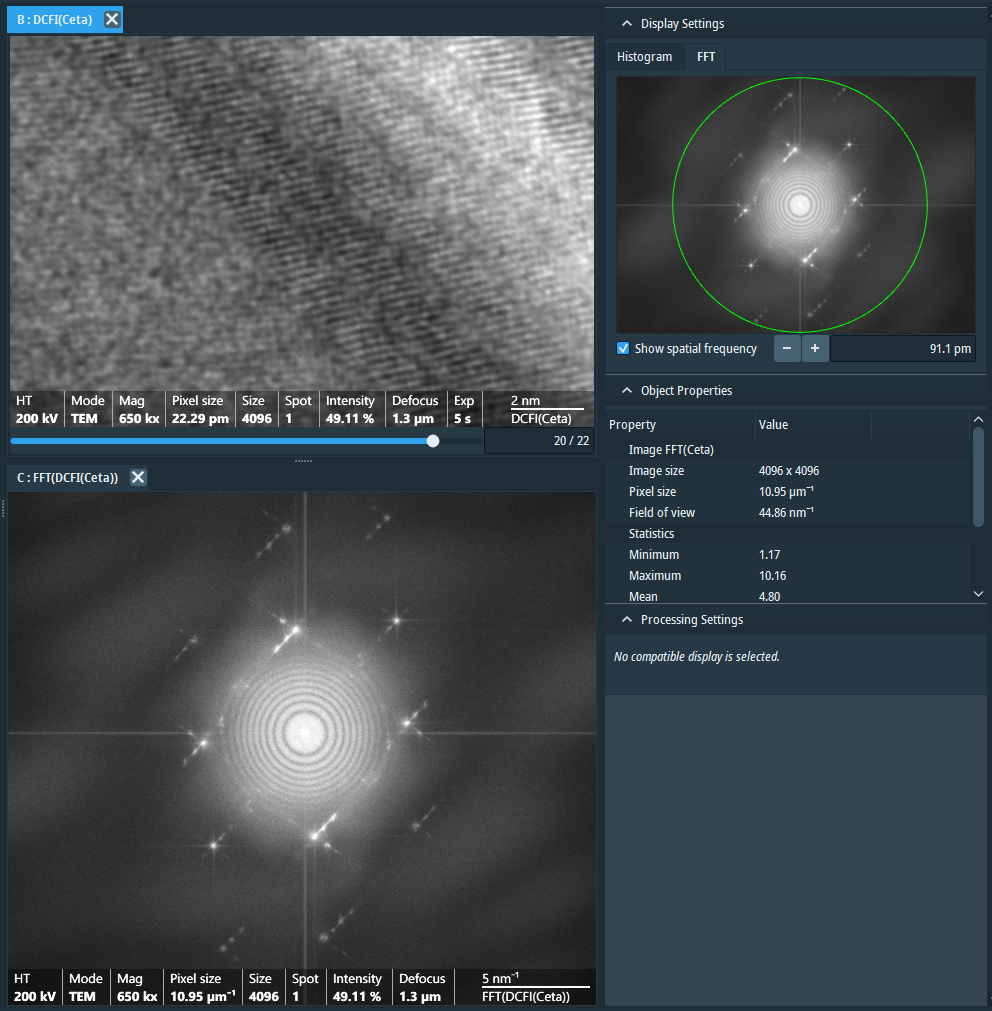
This is a transmission electron microscope equipped with the high sensitive, high resolution and high signal/noise (S/N) Ceta-S/F camera, the MAPS software, the Tomography software, the EPU software and the inspec3D software. This is a 200KV state of the art transmission electron microscope to study the fine 3D-ultrastructure of tissues, cells and single bio-molecules either at room temperature or in their native states in frozen tissue sections or (for the purified bio-molecules) in vitreous ice. - Gatan 626 single tilt cryo-transfer sample holder
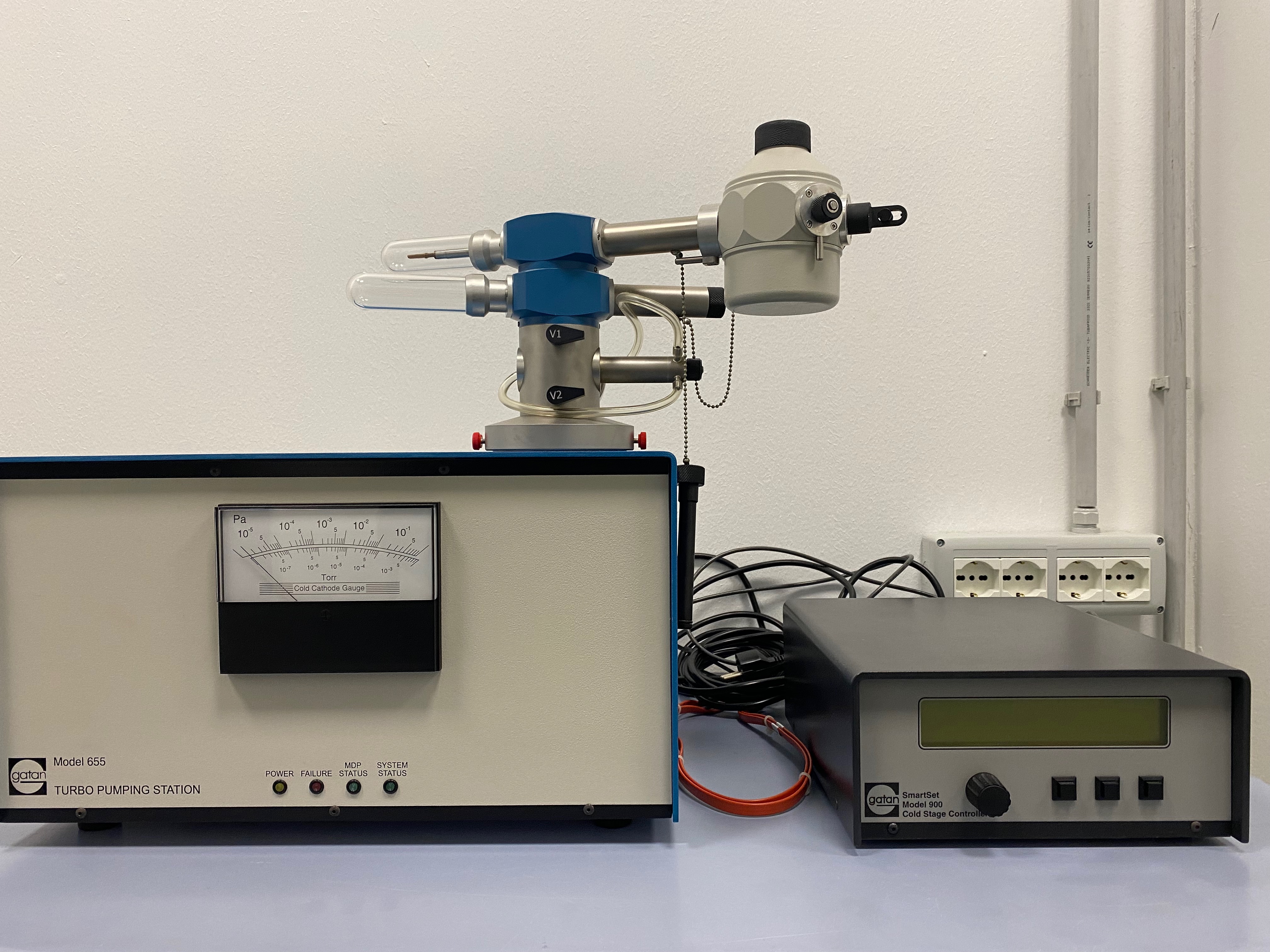
(Gatan 655 turbo molecular pumping station, Gatand 626 cryo-transfer-holder with temperature controller)
The complete cryo-transfer sample holder set-up includes: the Gatan 655 turbo molecular pumping station, the gatan cryo-transfer station and the Gatan 626 cryo-transfer- holder with the temperature controller unit. This is the essential device to load frozen bio-samples into the transmission electron microscope. This sample holder also allows cryo-electron tomography for the 3D reconstruction of tissue structures, cellular structures and single molecules 3D atomic structures in vitreous ice. - Fischione advanced tomography holder model 2020
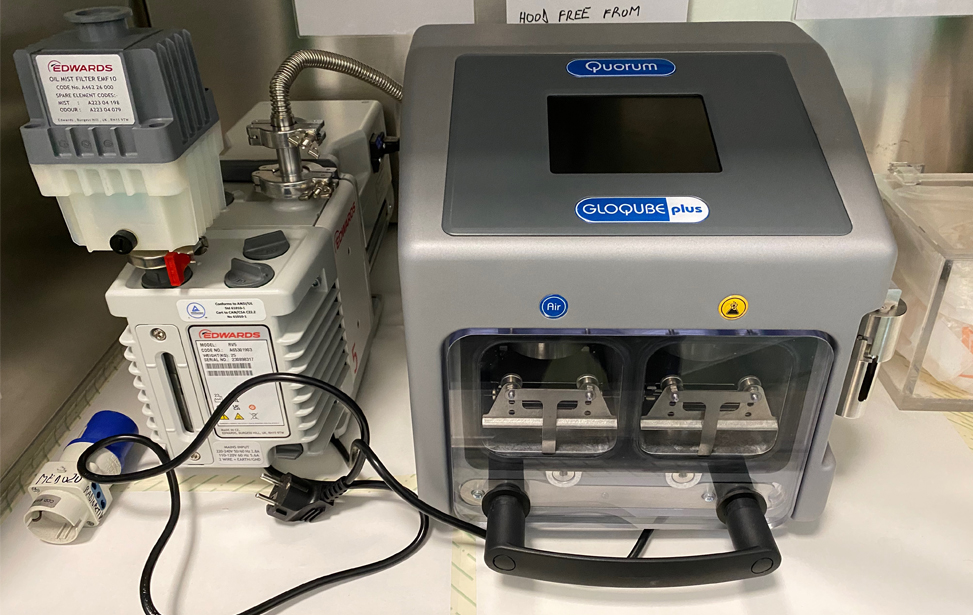
This the essential sample holder necessary for the acquisition of the tilted image series necessary for the 3D reconstruction of tomograms, which are instrumental for the studies of the 3D structure of the bio-samples at atomic resolution (either tissue structures, cells structures or the 3D structure of the bio-molecules) at room temperature. - Quorum- GloQube plus-glow discharger
This is the instrument necessary to treat the surface of the electron microscopy supports, which are necessary as support surfaces to apply the bio samples, which will be analyzed with all the high-resolution techniques adopted at the electron microscopy facility - FEI Tecnai 12 G2 biotwin transmission electron microscope
This is a transmission electron microscope with a side mounted Gatan Orius SC-1000 camera. This is an entry-level transmission electron microscope equipped with the open source serial EM software. It allows the acquisition of tomograms at room temperature with a dedicated room temperature tomography sample holder. It offers an extended field of view ideal for quick quality control of bio-sample, quick sample screenings and it is an ideal easy to use platform for the introductory electron microscopy trainings for beginners users from IFOM research groups that would like to start to approach the electron microscopy techniques in a more detailed fashion - Leica EM ACE600 electron beam (e-beam) metallic coater
This instrument is used to produce ultra-thin, high-quality metallic layers through the focused electron beam technology on a wide range of electron microcopy supports, ensuring optimal sample preparation for analysis. It is also essential for preparing rotary shadowing samples to visualize nucleic acids. The electron beam configuration (e-beam) is ideal to obtain high quality electron microscopy supports to study the atomic 3D structures of the single bio-molecules - Leica MED020 electron beam (e-beam) metallic coater
This instrument is used to produce ultra-thin, high-quality metallic layers (including carbon and platinum layers) on a wide range of electron microscopy supports, ensuring optimal sample preparation for analysis. It is also essential for preparing rotary shadowing samples to visualize nucleic acids.
Facility Acknowledgment
According to our Core Facilities Guidelines, if Facility services were used in work described in a publication, the Facility must be acknowledged, citing both the Facillity name and its Research Resource Identifier (RRID). A typically acknowledgement might look like:
The research described was supported by the IFOM Electron Microscopy Single Molecules Core Facility (RRID: SCR_026864)
Co‑Authorship
If a staff member of an IFOM or Cogentech Core Facility contributes substantially to a published work, they should be included as a co‑author.
More information is available in Recommended guidelines and Acknowledging and citing core facilities
Examples of substantial contributions that may justify co‑authorship include:
- Contribution to experimental design or optimization
- Development or significant adaptation of methods or protocols
- Advanced data analysis and interpretation
- Intellectual input influencing the scientific direction of the study
Routine technical support or standard service provision generally warrants acknowledgment rather than co‑authorship.
- Oxidative stress at telomeres triggers internal DNA loops, TRF1 dissociation, and TRF2-dependent R-loops
Nguyen TT, Mazzucco G, Kyriacou E, Lunardi T, Brandl L, Ahmed W, Doksani Y, Lingner J.
Nucleic Acids Res. 2025 Apr 10;53(7):gkaf285. doi: 10.1093/nar/gkaf285. PMID: 40219969 - HMCES corrupts replication fork stability during base excision repair in homologous recombination-deficient cells
Peña-Gómez MJ, Rodríguez-Martín Y, Del Rio Oliva M, Wijesekara Hanthi Y, Berrada S, Freire R, Masson JY, Reyes JC, Costanzo V, Rosado IV.
Sci Adv. 2025 Mar 28;11(13):eads3227. doi: 10.1126/sciadv.ads3227. Epub 2025 Mar 26.PMID: 40138423 - RAD51 protects abasic sites to prevent replication fork breakage
Hanthi YW, Ramirez-Otero MA, Appleby R, De Antoni A, Joudeh L, Sannino V, Waked S, Ardizzoia A, Barra V, Fachinetti D, Pellegrini L, Costanzo V. Mol Cell. 2024 Aug 22;84(16):3026-3043.e11. doi: 10.1016/j.molcel.2024.07.004. PMID: 39178838 Free article. - Unveiling the mechanistic link between extracellular amyloid fibrils, mechano-signaling and YAP activation in cancer.
Farris F, Elhagh A, Vigorito I, Alongi N, Pisati F, Giannattasio M, Casagrande F, Veghini L, Corbo V, Tripodo C, Di Napoli A, Matafora V, Bachi A. Cell Death Dis. 2024 Jan 11;15(1):28. doi: 10.1038/s41419-024-06424-z. PMID: 38199984 Free PMC article. - Sen1 and Rrm3 ensure permissive topological conditions for replication termination
Ramveer Choudhary, Joanna Niska-Blakie, Mohamood Adhil, Giordano Liberi, Yathish Jagadheesh Achar, Michele Giannattasio, Marco Foiani.
Cell Rep. 2023 Jul 4;42(7):112747. doi: 10.1016/j.celrep.2023.112747. - RFWD3 promotes ZRANB3 recruitment to regulate the remodeling of stalled replication forks.
Moore CE, Yalcindag SE, Czeladko H, Ravindranathan R, Wijesekara Hanthi Y, Levy JC, Sannino V, Schindler D, Ciccia A, Costanzo V, Elia AEH.
J Cell Biol. 2023 May 1;222(5):e202106022. doi: 10.1083/jcb.202106022. Epub 2023 Apr 10. PMID: 37036693 - The telomerase reverse transcriptase elongates reversed replication forks at telomeric repeats.
Huda A, Arakawa H, Mazzucco G, Galli M, Petrocelli V, Casola S, Chen L, Doksani Y.
Sci Adv. 2023 Mar 22;9(12):eadf2011. doi: 10.1126/sciadv.adf2011. Epub 2023 Mar 22. PMID: 36947627 Free PMC article. - Short-term molecular consequences of chromosome mis-segregation for genome stability
Lorenza Garribba , Giuseppina De Feudis , Valentino Martis , Martina Galli , Marie Dumont , Yonatan Eliezer , René Wardenaar , Marica Rosaria Ippolito, Divya Ramalingam Iyer, Andréa E Tijhuis , Diana C J Spierings , Michael Schubert , Silvia Taglietti , Chiara Soriani , Simon Gemble , Renata Basto , Nick Rhind , Floris Foijer , Uri Ben-David , Daniele Fachinetti , Ylli Doksani , Stefano Santaguida
Nat Commun . 2023 Mar 11;14(1):1353. doi: 10.1038/s41467-023-37095-7. - Enrichment of DNA replication intermediates by EdU pull down
Pessina, F., Romussi, A., Piccini, D., Mazzucco, G, Varasi, M., and Doksani, Y. (2022)
Methods in Cell Biology. Book Chapter - POLq prevents MRE11-NBS1-CtIP dependent fork breakage in the absence of BRCA2/RAD51 by filling lagging strand gaps.
Anjali Mann, Miguel Angel Ramirez-Otero, Anna De Antoni, Yodhara WijesekaraHanthi, Vincenzo Sannino, Giorgio Baldi, Lucia Falbo, Anna Schrempf, Sara Bernardo, Joanna Loizou, Vincenzo Costanzo.
Mol Cell 2022. Volume 82, Issue 22,2022,Pages 4218-4231.e8,ISSN 1097-2765, https://doi.org/10.1016/j.molcel.2022.09.013. - Rad51-mediated replication of damaged templates relies on monoSUMOylated DDK kinase
Chinnu Rose Joseph, Sabrina Dusi, Michele Giannattasio, Dana Branzei
Nat Commun 2022. May 5;13(1):2480. doi: 10.1038/s41467-022-30215-9. - Parental histone deposition on the replicated strands promotes error-free DNA damage tolerance and regulates drug resistance
Valeria Dolce, Sabrina Dusi, Michele Giannattasio, Chinnu Rose Joseph, Marco Fumasoni, Dana Branzei
Genes Dev 2022. Feb 1;36(3-4):167-179. doi: 10.1101/gad.349207.121. Epub 2022 Feb 3. - A rapid method to visualize human mitochondrial DNA replication through rotary shadowing and transmission electron microscopy
Martin Kosar, Daniele Piccini, Marco Foiani, Michele Giannattasio
Nucleic Acids Research, Volume 49, Issue 21, 2 December 2021, Page e121, https://doi.org/10.1093/nar/gkab770 - REV1-Polζ maintains the viability of homologous recombination-deficient cancer cells through mutagenic repair of PRIMPOL-dependent ssDNA gaps
Angelo Taglialatela, Giuseppe Leuzzi , Vincenzo Sannino , Raquel Cuella-Martin , Jen-Wei Huang , Foon Wu-Baer , Richard Baer , Vincenzo Costanzo , Alberto Ciccia.
Mol Cell. 2021 Oct 7;81(19):4008-4025.e7. doi: 10.1016/j.molcel.2021.08.016. - Checkpoint-mediated DNA polymerase ε exonuclease activity curbing counteracts resection-driven fork collapse.
Grazia Pellicanò, Mohammed Al Mamun, Dolores Jurado-Santiago , Sara Villa-Hernández , Xingyu Yin , Michele Giannattasio , Michael C Lanz , Marcus B Smolka , Joseph Yeeles , Katsuhiko Shirahige, Miguel García-Díaz , Rodrigo Bermejo
Mol Cell. 2021 Jul 1;81(13):2778-2792.e4. doi: 10.1016/j.molcel.2021.04.006. Epub 2021 Apr 30 - Smc5/6 functions with Sgs1-Top3-Rmi1 to complete chromosome replication at natural pause sites.
Sumedha Agashe , Chinnu Rose Joseph , Teresa Anne Clarisse Reyes , Demis Menolfi , Michele Giannattasio , Anja Waizenegger , Barnabas Szakal , Dana Branzei
Nat Commun. 2021 Apr 8;12(1):2111. doi: 10.1038/s41467-021-22217-w. - The human nucleoporin Tpr protects cells from RNA-mediated replication stress.
Martin Kosar , Michele Giannattasio , Daniele Piccini , Apolinar Maya-Mendoza , Francisco García-Benítez , Jirina Bartkova , Sonia I Barroso , Hélène Gaillard , Emanuele Martini, Umberto Restuccia , Miguel Angel Ramirez-Otero, Massimiliano Garre, Eleonora Verga, Miguel Andújar-Sánchez , Scott Maynard , Zdenek Hodny , Vincenzo Costanzo , Amit Kumar , Angela Bachi , Andrés Aguilera , Jiri Bartek , Marco Foiani
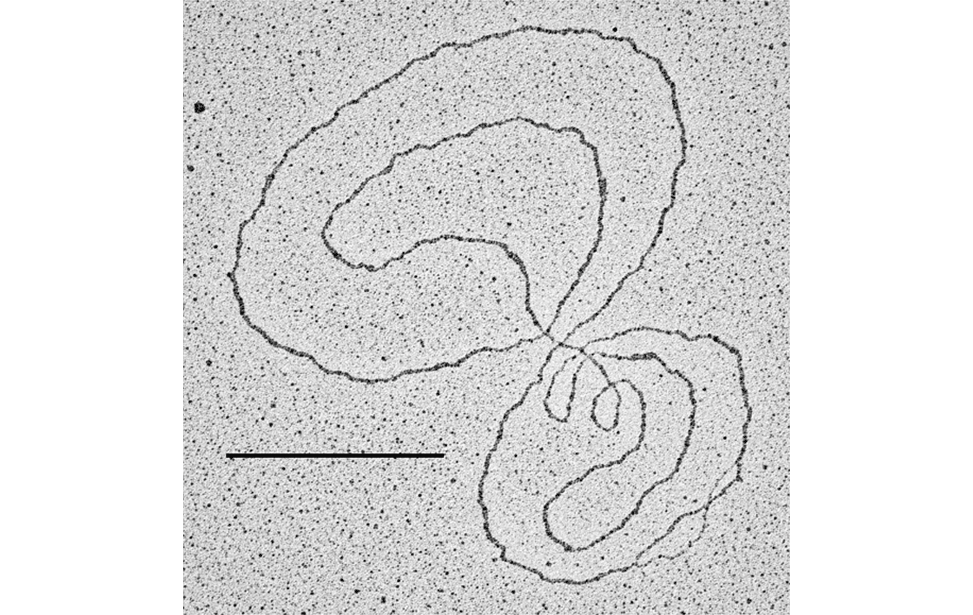
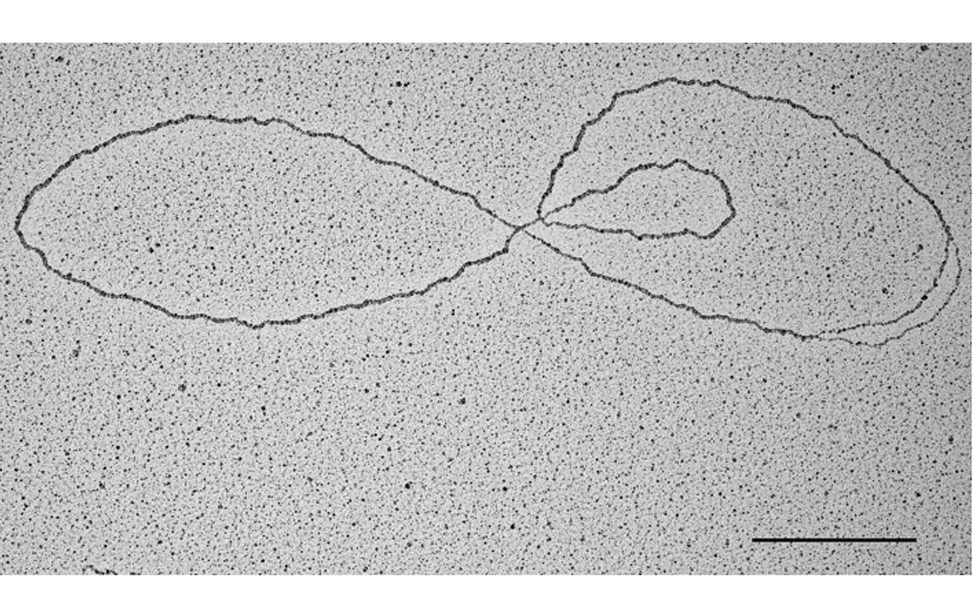
Nat Commun.2021 Jun 24;12(1):3937. doi: 10.1038/s41467-021-24224-3. - Telomere damage induces internal loops that generate telomeric circles.
Giulia Mazzucco, Armela Huda, Martina Galli, Daniele Piccini, Michele Giannattasio, Fabio Pessina, Ylli Doksani
Nat Commun. 2020 Oct 20;11(1):5297. doi: 10.1038/s41467-020-19139-4. - Physiological Tolerance to ssDNA Enables Strand Uncoupling during DNA Replication.
Amaia Ercilla, Jan Benada , Sampath Amitash, Gijs Zonderland, Giorgio Baldi , Kumar Somyajit, Fena Ochs ,Vincenzo Costanzo , Jiri Lukas ,Luis Toledo
Cell Rep 2020 Feb 18;30(7):2416-2429. doi: 10.1016/j.celrep.2020.01.067
- Dna2 processes behind the fork long ssDNA flaps generated by Pif1 and replication-dependent strand displacement.
Rossi SE, Foiani M, Giannattasio M.
Nature Communications. 2018 Nov 16;9(1):4830. doi: 10.1038/s41467-018-07378-5. - AND-1 fork protection function prevents fork resection and is essential for proliferation.
Abe T, Kawasumi R, Giannattasio M, Dusi S, Yoshimoto Y, Miyata K, Umemura K, Hirota K, Branzei D.
Nature Communications. 2018 Aug 6;9(1):3091. doi: 10.1038/s41467-018-05586-7. - Restoration of Replication Fork Stability in BRCA1- and BRCA2-Deficient Cells by Inactivation of SNF2-Family Fork Remodelers.
Taglialatela A, Alvarez S, Leuzzi G, Sannino V, Ranjha L, Huang JW, Madubata C, Anand R, Levy B, Rabadan R, Cejka P, Costanzo V, Ciccia A.
Molecular Cell. 2017 Oct 19;68(2):414-430.e8. doi: 10.1016/j.molcel.2017.09.036. - Fanconi-Anemia-Associated Mutations Destabilize RAD51 Filaments and Impair Replication Fork Protection.
Karina Zadorozhny, Vincenzo Sannino, Ondrej Beláň, Jarmila Mlčoušková, Mário Špírek, Vincenzo Costanzo, Lumír Krejčí.
Cell Rep 2017 Oct 10;21(2):333-340. doi: 10.1016/j.celrep. 2017.09.062. - Smarcal1-Mediated Fork Reversal Triggers Mre11-Dependent Degradation of Nascent DNA in the Absence of Brca2 and Stable Rad51 Nucleofilaments.
Kolinjivadi AM, Sannino V, De Antoni A, Zadorozhny K, Kilkenny M, Técher H, Baldi G, Shen R, Ciccia A, Pellegrini L, Krejci L, Costanzo V.
Molecular Cell. 2017 Sep 7;67(5):867-881.e7. doi: 10.1016/j.molcel.2017.07.001. Epub 2017 Jul 27. - Xenopus laevis as Model System to Study DNA Damage Response and Replication Fork Stability
Vincenzo Sannino , Federica Pezzimenti , Stefania Bertora , Vincenzo Costanzo. (2017)
Methods Enzymol. 2017;591:211-232. doi: 10.1016/bs.mie.2017.03.018. Epub 2017 Apr 8. - Centromeric DNA replication reconstitution reveals DNA loops and ATR checkpoint suppression.
Aze A, Sannino V, Soffientini P, Bachi A, Costanzo V.
Nature Cellular Biology. 2016 Jun;18(6):684-91. doi: 10.1038/ncb3344. Epub 2016 Apr 25. - Rad53-Mediated Regulation of Rrm3 and Pif1 DNA Helicases Contributes to Prevention of Aberrant Fork Transitions under Replication Stress.
Rossi SE, Ajazi A, Carotenuto W, Foiani M, Giannattasio M.
Cell Reports. 2015 Oct 6;13(1):80-92. doi: 10.1016/j.celrep.2015.08.073. Epub 2015 Sep 24. - Error-free DNA damage tolerance and sister chromatid proximity during DNA replication rely on the Polα/Primase/Ctf4 Complex.
Fumasoni M, Zwicky K, Vanoli F, Lopes M, Branzei D.
Molecular Cell. 2015 Mar 5;57(5):812-823. doi: 10.1016/j.molcel.2014.12.038. Epub 2015 Feb 5. - Visualization of recombination-mediated damage bypass by template switching.
Giannattasio M, Zwicky K, Follonier C, Foiani M, Lopes M, Branzei D.
Nature Structural & Molecular Biology. 2014 Oct;21(10):884-92. doi: 10.1038/nsmb.2888. Epub 2014 Sep 7.
DNA
Proteins
Equipment

Michele Giannattasio
Michele Giannattasio is a molecular biologist and geneticist with extensive expertise in transmission electron microscopy (TEM) for the structural analysis of DNA, RNA molecules and proteins. He obtained his Master’s degree in Molecular Biology (1998) and PhD in Genetics (2002) from the University of Milan. Michele is officer-level staff scientific and technological sector at the University of Milano (department of oncology and haematology-oncology) Michele’s interest in TEM began in 2008, when he used this technique to visualize long single-stranded DNA (ssDNA) gaps generated during Nucleotide Excision Repair (NER). In 2010, he joined IFOM as a staff scientist, applying and developing TEM-based methods for the visualization and analysis of chromosomal DNA intermediates. Over the years, he has worked closely with multiple IFOM research groups, contributing to the study of DNA damage tolerance, DNA replication, DNA recombination and DNA repair mechanisms. His international experience includes research at the Electron Microscopy Research Centre of the university of Zurich (centre for microscopy and image analysis) and at the Institute of Molecular Cancer Research in Zurich, where he specialized in advanced TEM methodologies and sample preparation techniques. In 2017, Michele became Head of the Electron Microscopy Single Molecule Core Facility at IFOM, where he continues to expand the application of TEM in genome and protein research, making electron microscopy based ultrastructural approaches available to the IFOM scientific community.