Program Genome Diagnostics
Background and current state-of-the-art
The advent of massive parallel DNA sequencing has revolutionized the biomedical research field ensuring rapid and affordable deciphering of the human genome sequence. This has led to a dramatic acceleration in the identification of genetic alterations responsible for several congenital and acquired diseases. In particular, taking advantage of next generation sequencing (NGS) technology, both national and international large-scale genome sequencing projects have made substantial progress in the reconstruction of the spectrum of recurrent genetic alterations present in sporadic and inherited forms of cancer.
The continuous accumulation of genome sequencing information from primary cancers worldwide has witnessed a parallel boosting of genome bioinformatics. The development (and continuous improvement) of dedicated computer algorithms allows nowadays an increasingly faster analysis of genome data, ensuring the concomitant investigation of multiple aspects of cancer biology. Such information has already led to a substantial revisiting of the basic concepts underlying cancer initiation, progression and metastasis.
Large-scale genome/exome sequencing studies have also accelerated the identification of new genetic factors that confer inherited predisposition to develop cancer. This knowledge is rapidly expanding and is expected to revolutionize in the next 10 years the healthcare system in many countries worldwide. NGS technology offers already the opportunity to perform cost-affordable individual, family- or population-based screenings for a growing list of cancer susceptibility genes. This information will substantially improve the identification of individuals at risk of developing cancer, providing thereby a critical support in the decision making process of the risk reducing or surveillance options.
Finally, the implementation of NGS technology applied to cancer genomes has represented a turning point for medical treatment decisions. Indeed, the deciphering of the mutation spectrum associated with individual primary cancers offers the opportunity to design personalized cancer treatments. NGS also allows an accurate estimate of tumor cell heterogeneity affecting individual cancers. The latter information is relevant to explain the bases for primary and acquired tumor resistance to current therapies. Revealing the genetic complexity of individual cancers through NGS-based genetic screenings is expected to provide the oncologist with the possibility to design patient-tailored combinatorial treatments targeting concomitantly multiple independent tumor subpopulations composing the cancer.
Our Goals
IFOM investigators are committed to study the genetic factors predisposing to cancer and the mechanisms promoting cancer development and metastasis, looking at the “cancer problem” from a genetic, molecular, cellular and organismal perspective. In IFOM, studies on pre-clinical tumor models are coupled to investigations on human primary cancers and on cancer-prone families, in close collaboration with oncologists, pathologist, surgeons and genetic counselors from national and international centers.
We have a particular interest in translating findings from preclinical cancer models and cancer genome sequencing efforts into the design of novel genetic tests for screening purposes. We also intend to widen the portfolio of genetic variants to be screened to establish an inherited increased risk to develop particular forms of cancer. The validation stage will be followed by the implementation of such tests in the clinics where tailored treatments could be offered to carriers of specific mutations.
Moreover, IFOM investigators are actively engaged in investigations on chromosome and genome biology to understand how several different types of genetic alterations that are recurrently observed in cancer cells contribute to its predisposition, development and metastatic evolution.
Finally, a deeper understanding of the genetic bases responsible for cancer predisposition, initiation, development and progression is combined in IFOM with bioinformatics analyses of publicly accessible human (cancer) genome databases, to ultimately design novel genetic tests and genome-focused methodologies intended to help anticipate cancer diagnosis and guide its treatment.
The IFOM Genome Diagnostics Program (GDP)
This program merges the unique scientific environment of IFOM with advanced genomic technologies, to develop genetic tests and methodologies for cancer prevention, diagnosis and prognosis and to guide personalized treatment. Within the GDP, research activities are channeled into two major areas of investigations:
1) Cancer diagnosis and treatment of cancer diseases.
Active collaborations with pathologists, oncologists and surgeons allow IFOM investigators to conduct a systematic analysis of candidate gene alterations associated with:
- a. Specific cancer subtypes.
- b. Resistance/response to standard therapies.
- c. Subclonal distribution of tumor cells.
- d. Efficacy of novel molecular- and/or immunotherapies
Through collaborations with oncologists, the mutational landscape of individual cancers is associated to the tumor histotype and to the clinical records of the patient, including present and past therapies. The matching of the genetic (e.g. gene mutation spectrum) and clinical data is expected to help clinicians to best stratify cancer patients who may benefit from specific therapeutic regimens, while preventing toxic side-effects to those patients whose cancer has the “wrong” genetic profile.
A substantial effort is also made to reconstruct the clonal composition of primary cancers. In this context, the GDP is actively working to optimize methodologies enabling the molecular profiling of thousands of individual cancer cells from the same tumor, to obtain a representative genetic map of the subgroups of cancer cells composing the tumor bulk. This information will provide the molecular framework to design combo treatments targeting at the same time individual tumoral subpopulations composing the cancer mass.
2) Identification of cancer predisposing genetic factors.
In the last ten years, IFOM researchers have established collaborations with epidemiologists, geneticists, and molecular biologists from national and international institutions and are actively collaborating to projects carried on by large Consortia and aimed at:
- a. Identification of novel cancer predisposing genes and variants.
- b. Classification of variants of uncertain significance (VUS).
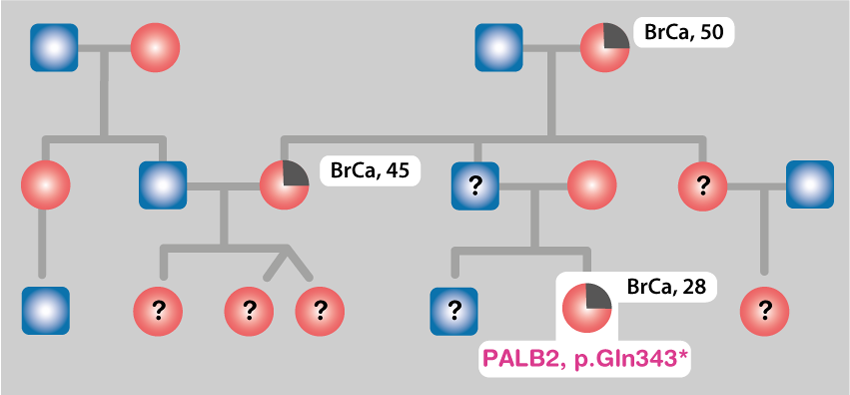
Mutations in BRCA1, BRCA2 and PALB2 genes, on one side, and in the genes of the mismatch repair system, on the other, confer a high risk to develop breast and colorectal cancer, respectively. Other genes and rare variants associated with the risk of developing these diseases are expected to exist. Their identification will increase the possibility to detect in the families and in the general population the individuals at risk for these (and possibly others) diseases.
Many of the detected variants in BRCA1, BRCA2, PALB2 and other breast cancer genes are associated with risk for the disease; however, some are currently unclassified and called VUS (variants of uncertain significance). We are carrying on population- and family-based studies and functional analysis aimed at classifying many of these variants clarifying which are risk factors and which are neutral. This will allow providing more precise risk estimates in a larger fraction of individuals from families and general population.
Key resources mobilized
To achieve these goals, GDP relies on a solid technological infrastructure centered on state-of-the-art NGS technology, which is coordinated by the IFOM genomic unit. Also, crucial for the success of the program are trained bioinformaticians involved in the analysis, interpretation and storage of the an enormous amount of data derived from the sequencing of thousands of exomes and genomes.
The rapid transfer of the knowledge from the bench to the bedside will be ensured by the close collaboration of GDP investigators with the Cogentech Genetic Tests (GT) Unit, which has served in the past 10 years local hospitals performing diagnostic genetic tests accredited by the National Health System for breast and colon cancer predisposing genes. Novel genetic tests developed through GDP will be made available to clinicians through the Cogentech GT Unit, which will contribute to the optimization of the procedures to deliver in a time- and cost-affordable fashion the tests to the clinicians and genetic counselors.
Expected impact
The GDP is a research-based program that exploits the knowledge from advanced cancer genome investigations and basic studies on preclinical cancer models, cancer patients and cancer-prone families, to improve cancer diagnosis and treatment and to identify new tumor predisposing genes. We expect to develop cost-affordable genetic tests to improve cancer prevention, and its early detection, and to personalize cancer treatment. The consolidation of collaborations between IFOM investigators and oncologists, pathologists, surgeons and genetic counselors is instrumental for the rapid validation of the newly designed genetic tests in the clinical setting. Research conducted within IGDRP is expected to help a broad spectrum of cancer patients, and individuals at risk of developing the disease. We believe that a larger fraction of such individuals should benefit from risk reducing options, improved strategies of surveillance for early disease detection, and better clinical treatments.