Genetics of B Cells and Lymphomas
Research Project
The pool of peripheral B lymphocytes is established through an ordered set of molecular events initiated in the bone marrow with the assembly on the surface of the cells of a functional, non-autoimmune, B cell antigen receptor (BCR) complex. Expression and signaling through the BCR is essential for developing B cells to differentiate respectively into follicular, marginal zone or B-1 B cells, each of these subsets displaying a specific effector function. Basal signaling through the BCR is required for survival of resting mature B cells, which in turn become rapidly activated and undergo proliferation upon BCR-mediated antigen recognition. Signals arising from BCR engagement leading to B cell proliferation, survival and differentiation are commonly integrated by others initiated from receptors such as BAFF-R, CD40 and the family of Toll-like receptors recognizing a variety of microbial antigens.
The germinal center reactionUpon recognition of foreign antigens through the BCR, in the presence of T-cell help, B cells are recruited into germinal centers (GCs) formed in spleen and lymph nodes during T-cell dependent antibody responses. During the GC reaction, B cells undergo an initial phase of proliferation followed by cell-cycle arrest and selection based on the affinity of the BCR for the cognate antigen. The establishment of a productive antibody response to a foreign antigen depends on the selection within the GC of a limited pool of B cells expressing high affinity antigen receptors generated in situ through the process of immunoglobulin (Ig) somatic mutation. GC B cells also undergo replacement of the constant region of the Ig heavy chains through the process of Ig class-switch recombination, thereby changing the effector functions of the BCR. Finally, high-affinity, isotype-switched, B cells exit the GC to become long-lived memory B cells or antibody-producing plasma cells.
The germinal center origin of B cell lymphomasThe majority of human B cell malignancies arise from mature B cells recruited into the GC reaction. GC B cell lymphomas are classified on the basis of histological appearance and molecular features as Hodgkin and non-Hodgkin B cell lymphomas. The latter group includes several tumor types such as Burkitt lymphoma (BL), follicular lymphoma (FL), diffuse-large B cell lymphomas (DLBCL) and lymphomas affecting mucosa-associated lymphoid tissues (MALT). Despite the common GC B-cell origin, Hodgkin and the different forms of non-Hodgkin B cell lymphomas differ from each other by the involvement of different oncogenic pathways driving the tumorigenic process.
Conditional gene targeting as a tool to study germinal center B cell biologyTo study gene function in GC B cells we employ the Cre/loxP recombination system to generate conditional gene-targeted mice. Specifically, transgenic animals are generated from embryonic stem cells manipulated in vitro through DNA homologous recombination. Experimental mice obtained with this procedure carry, on the one hand, a transgene coding for the Cre recombinase expressed in a cell-type and stage-specific manner (e.g. GC B cells) and, on the other, the gene of interest flanked by loxP sites. In this way, Cre-mediated recombination leads to inducible loss of gene function in a cell-type and stage-specific manner. We employ the Cre/loxP recombination system also to induce in vivo activation of gene expression (e.g. proto-oncogenes) in a cell-type, stage-specific and temporal manner. To study gene function in GC B cells we dispose of Cϒ1-cre knock-in mice, allowing efficient Cre-mediated recombination in GC and post-GC B cells.
Objectives
Our group has three long-term goals:
- To dissect at the molecular level the key events that guide the transition of B cells through the sequential stages of the GC reaction. For this purpose, a series of conditional mouse mutant strains in which deletion of genes coding for a defined set of transcription factors, signaling proteins, surface receptors and chromatin remodeling enzymes will be specifically induced in GC B cells. The role of these factors in the control of GC B cell survival, selection and terminal differentiation will be determined. Also, their involvement in regulating Ig somatic mutation and class-switch recombination will be investigated. A similar approach will be followed to dissect the role of a defined set of microRNAs in GC B cell responses.
- To identify key genetic determinants controlling GC B-cell lymphoma initiation, maintenance and progression. Using existing mouse models of different subtypes of non-Hodgkin B cell lymphomas, we will delete in a conditional manner in lymphoma cells, at different stages of tumorigenesis, genes thought to play a critical role in (GC) B cell survival, proliferation and differentiation. The latter approach will be complemented by studies in which primary lymphoma cells of the different tumor subtypes will be transplanted into syngeneic recipient after infection with a library of lentiviral vectors expressing in an inducible manner short interfering RNAs directed against genes (initially B-cell specific) controlling specific functions of GC B cells. A major effort will be given to study the role of the BCR and its downstream signaling mediators in tumor B cell survival/proliferation. The final goal of these studies is to identify new therapeutic targets for the treatment of the different forms of GC B-cell malignancies.
- Tumor initiation, maintenance and progression results from functional cooperation between two or more oncogenes, often associated with loss of tumor suppressor gene activity. In this context, our group aims to identify partners of oncogenes whose function is commonly deregulated in specific forms of non-Hodgkin lymphomas including BL, FL and DLBCL. For this purpose, in mouse models of non-Hodgkin B cell lymphomas expressing the dominant oncogene as a transgene, insertional mutagenesis will be specifically induced in GC B cells. Acceleration of lymphoma onset in lymphoma-prone mice will confirm functional cooperation between oncogenes, eventually associated with loss of tumor suppressor gene function. The genes cooperating with the "primary" oncogene to accelerate disease will be cloned with PCR-based methods from genomic DNA extracted from a series of independent primary lymphomas. The biological relevance of the newly identified gene products will be validated analyzing their expression andactivation status in a series of primary human lymphoma specimens representative of the different types of non-Hodgkin B cell lymphomas. A similar approach will be attempted to identify oncogenic partners of the Epstein Barr Virus (EBV)-encoded proteins LMP1 and LMP2A expressed in the Hodgkin-Reed Sternberg tumor cells of EBV+ cases of classical Hodgkin lymphoma. For this purpose we will use mice, available in the lab, which express the viral proteins in a conditional manner.
04.02.2016 - rel 01
Photogallery
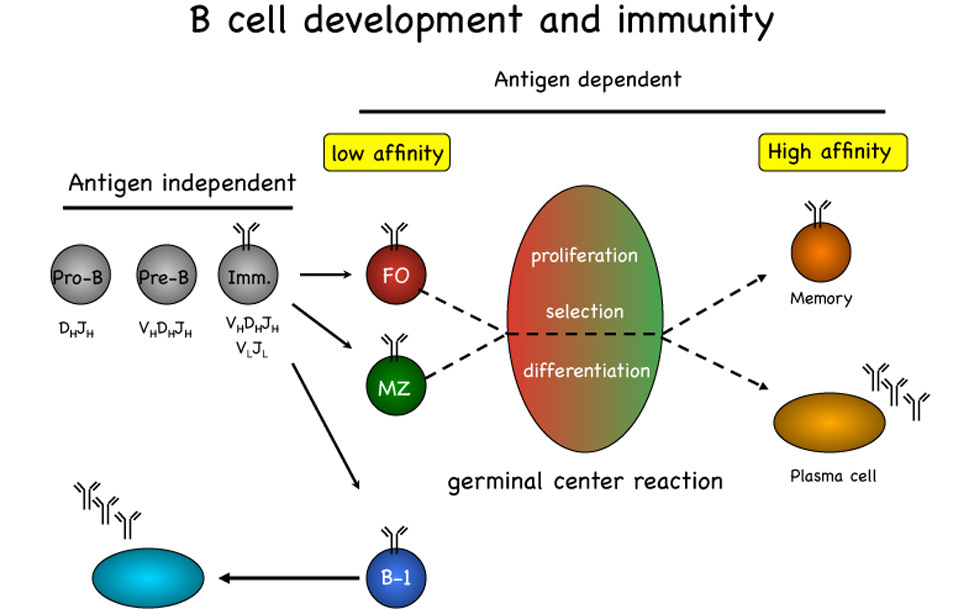 B-cell-development
B-cell-development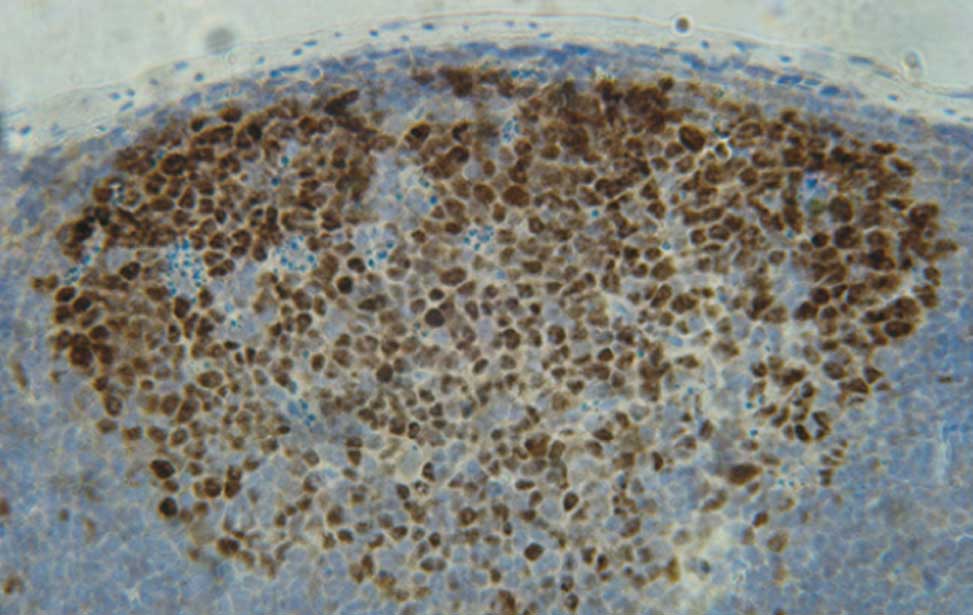 The germinal reaction
The germinal reaction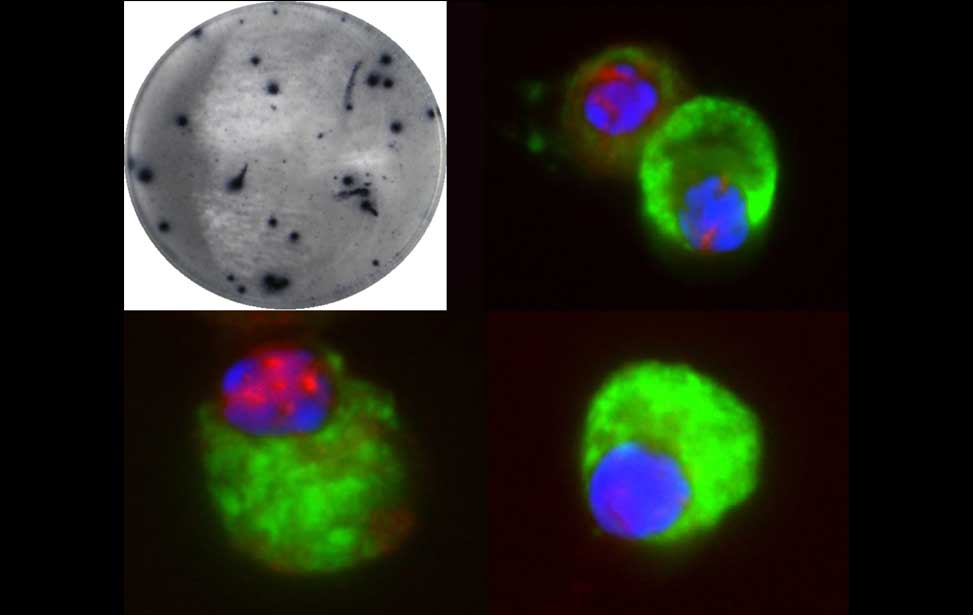 Antibody producing cells
Antibody producing cells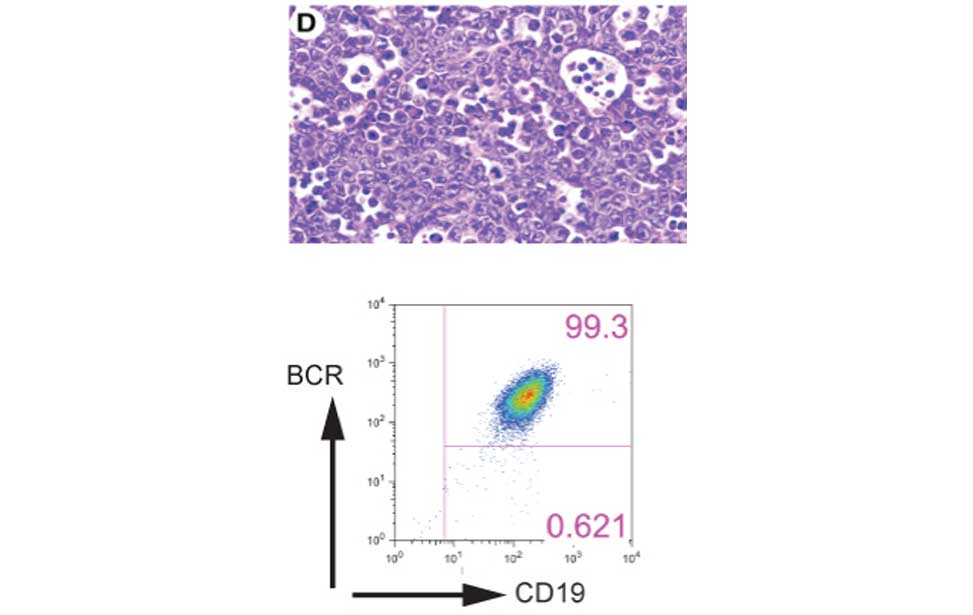 Burkitt lymphoma
Burkitt lymphoma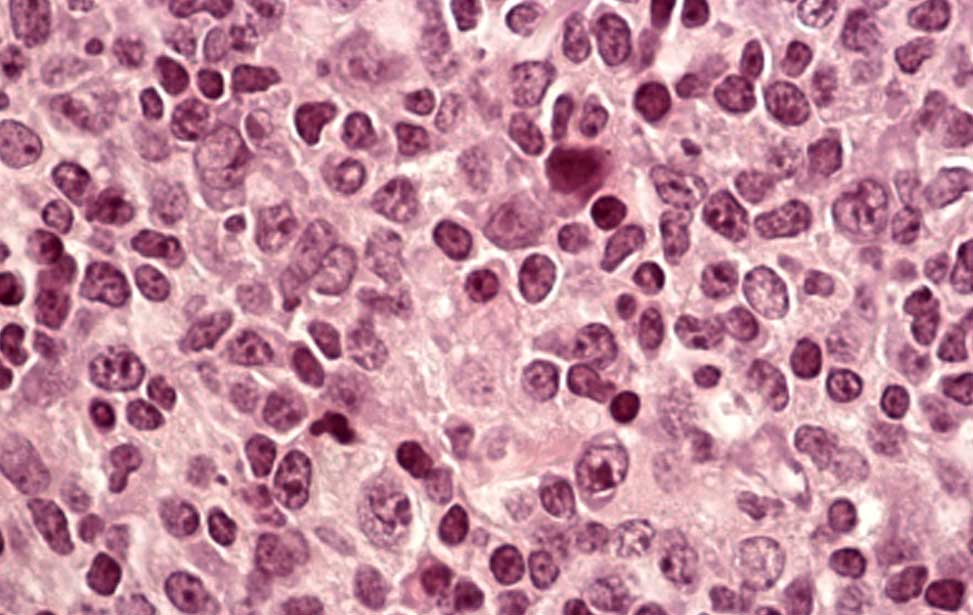 Diffuse Large B-cell lymphoma
Diffuse Large B-cell lymphoma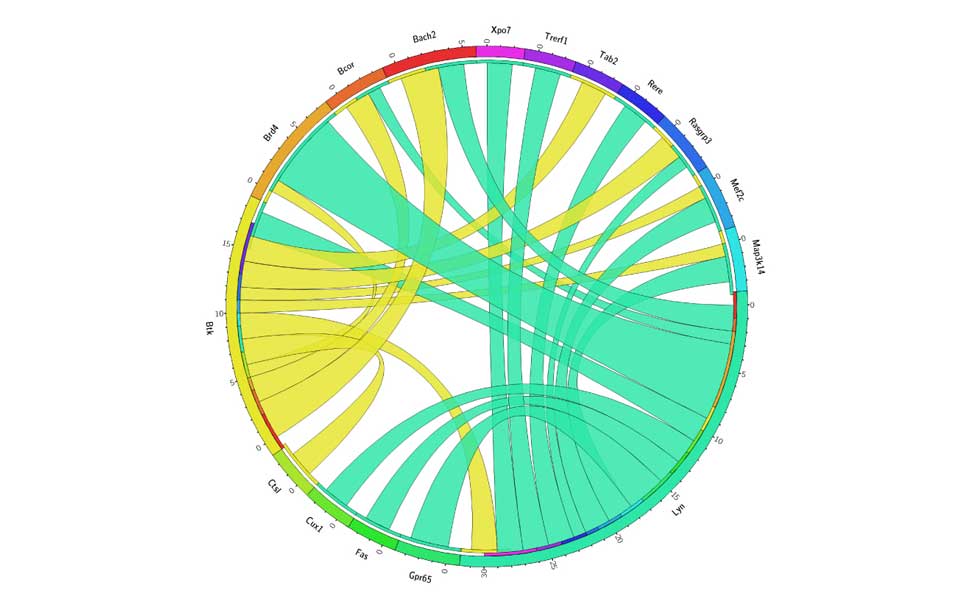 Epigenetics of B-lymphocytes and lymphomas
Epigenetics of B-lymphocytes and lymphomas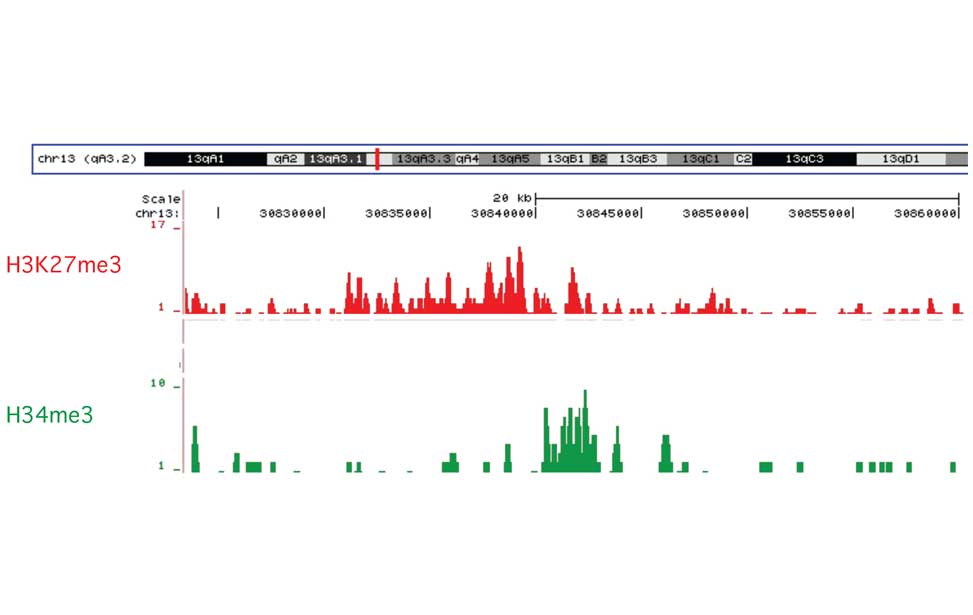 Screening for cancer mutations
Screening for cancer mutations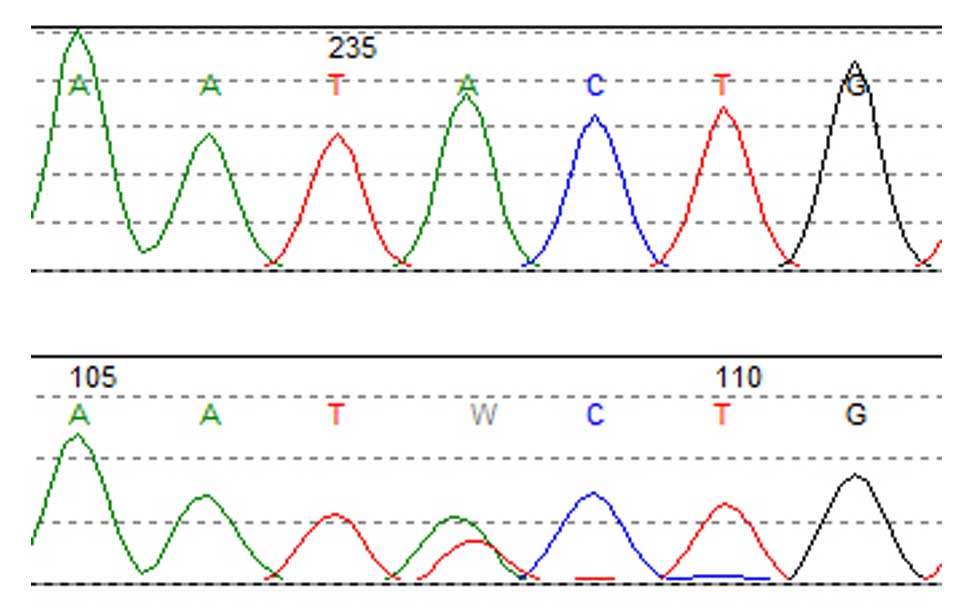 Genetics of Diffuse Large B-cell lymphomas
Genetics of Diffuse Large B-cell lymphomas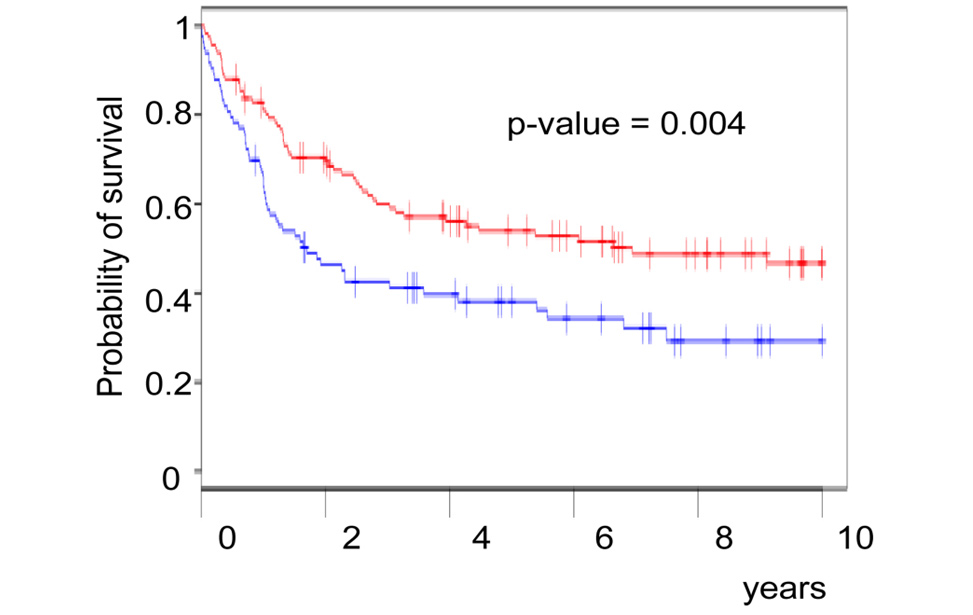 Novel prognostic factors in Diffuse Large B-cell lymphomas
Novel prognostic factors in Diffuse Large B-cell lymphomas 